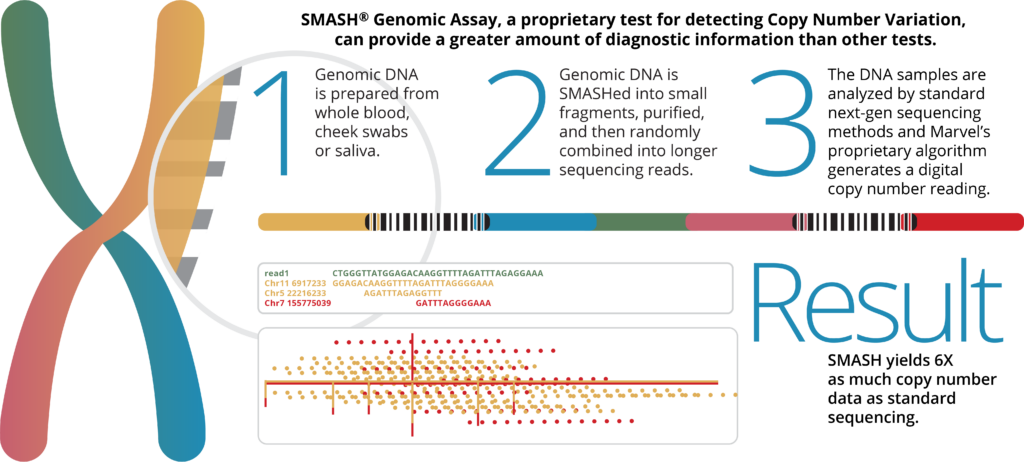
SMASH™ Genomic Assay
Research applications to accelerate target discovery
and drug development.
Marvel Genomics® has developed the SMASH Genomic Assay, a Copy Number Variation test that utilizes next-generation sequencing technology and proprietary software. There is great potential for genome sequencing to enhance patient care through improved diagnostic sensitivity and more precise therapeutic targeting. State-of-the-art DNA sequencing technologies and analytical algorithms can be adapted to fit clinical needs. We have focused on optimizing algorithms, giving particular attention to metrics such as reproducibility and accuracy, and standardizing genetic variant calls and interpretation.
Our proprietary approach simplifies DNA sequencing methods and analysis to rapidly identify genetic variants throughout the genome. We have validated samples using SMASH with higher sensitivity and specificity than existing approaches, giving us enhanced capabilities to identify the genetic causes of disease and develop more effective therapeutic molecules.
References:
1) SMASH method, Wang, Ronemus et al., Genome Research 844–851 (2016).
2) Nature Reviews Genetics, 507–522, 2016.
Clinical Issue / Unmet Needs
Copy number variants, which represent a large component of structural variation in the human genome, consist of duplication or deletion of DNA regions ranging from a few thousand to several million DNA bases in size. CNVs underlie a significant portion of the genetic factors causing autism spectrum disorders (ASD) – work for which Drs. Wigler, Ronemus and colleagues have been at the forefront for over a decade. CNVs have also been shown to play a role in many sporadic disorders such as congenital heart disease, cancer and schizophrenia as well as in patient responses to a variety of therapies.